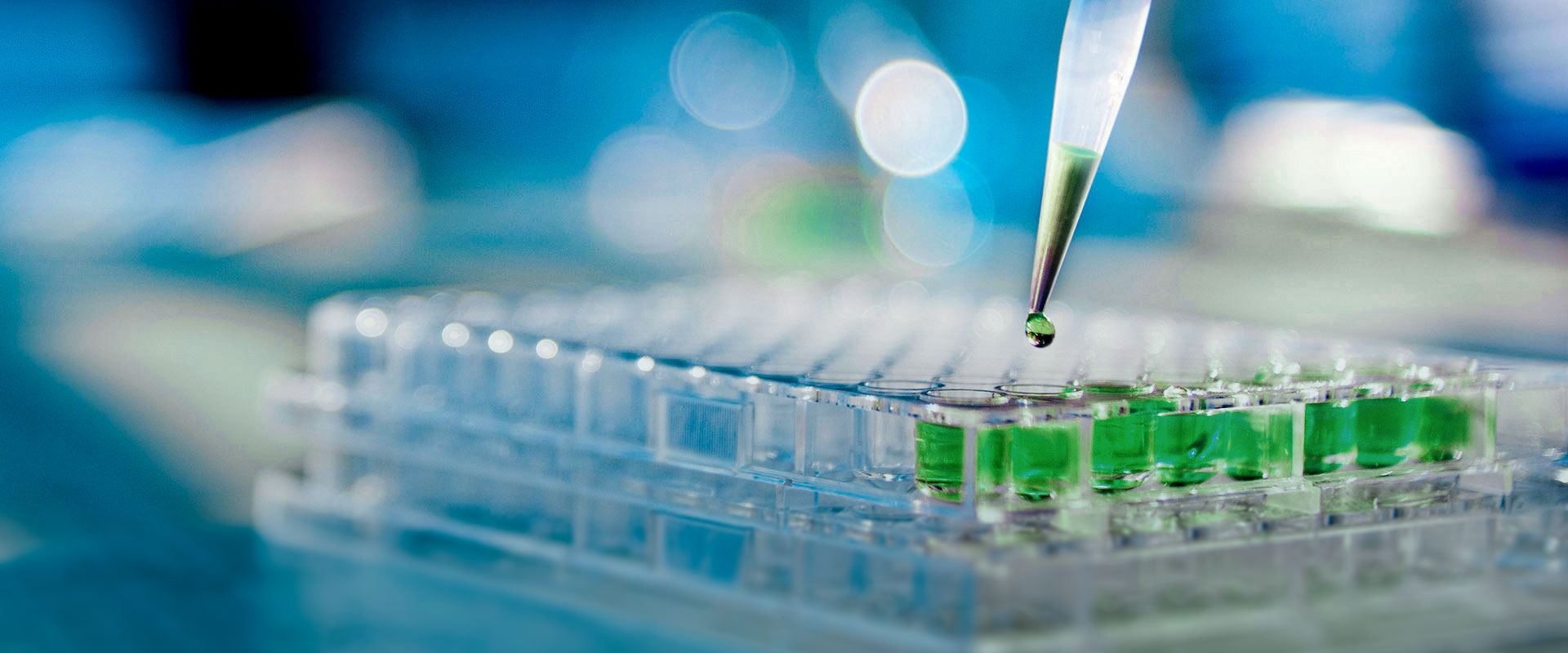
| Gene | Alteration |
| BRCA1 | SNV, InDel, LR |
| BRCA2 | SNV, InDel, LR |
The AmoyDx® BRCA Pro Panel comprehensively covers all protein coding regions, intron/exon boundaries, select introns, and UTR regions of the BRCA1 and BRCA2 genes. This panel enables the detection of germline/somatic SNVs/Indels as well as germline large rearrangements (LRs) from either whole blood or FFPE tissue samples.

Specifications
0.3 Gb for somatic variants
Technology
The AmoyDx HANDLE® technology is a proprietary amplicon-based method for NGS library preparation.
It offers a time-saving and cost-effective protocol that can be completed within 5 hours, requiring just about 1 hour of hands-on time.
During the library construction process, each individual DNA molecule is tagged with a unique molecular index (UMI) at both ends. This feature enables high sensitivity in variant detection by eliminating any library amplification and sequencing bias.
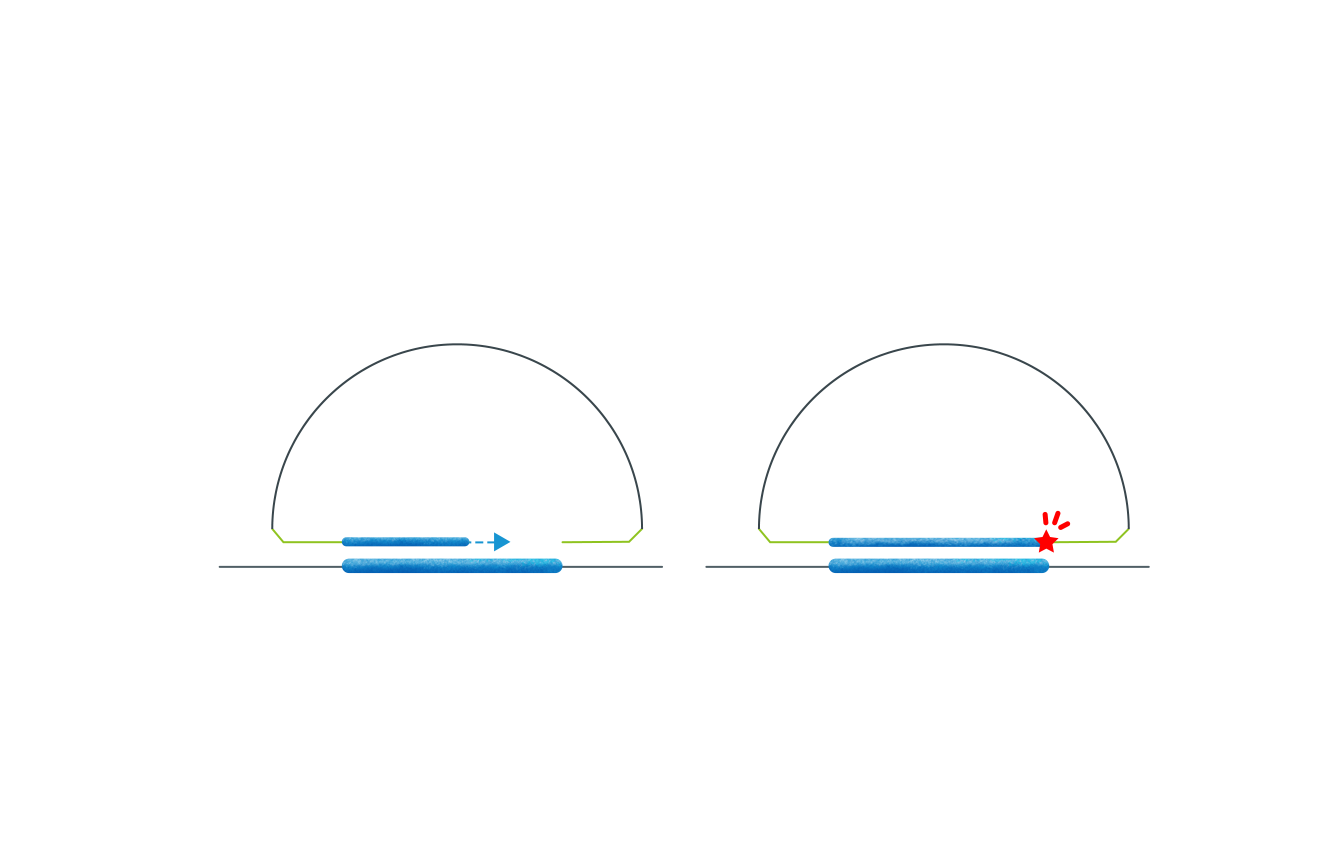
The probe consists of an extension arm and a ligation arm, both complementary to the target gene region.
Target-specific probes simultaneously hybridize to the DNA/cDNA fragment, forming a circular structure with the intended target captured between the probes. Subsequently, remaining linear probes, single-strand, and double-strand DNA are digested, leaving only the target circular DNA for PCR amplification to form the final library.
Sequencing data generated with AmoyDx NGS products can be conveniently and securely analyzed using the AmoyDx NGS Data Analysis System (ANDAS). The ANDAS Data Analyzer consists of a local server and preinstalled ANDAS software equipped with specific analysis modules tailored for analyzing data from various AmoyDx NGS products.
The ANDAS server is installed within the same local network as the sequencer, providing the laboratory with complete control over data and security.
This ensures the protection of valuable sequencing data and downstream analysis results, including sensitive patient information. Accessible via a computer within the network, the ANDAS server and its preinstalled software offer an intuitive user interface for swift and straightforward analysis generation.
Publications


1. Dong, Z., Wang, Y., Zhang, J. et al. Analyzing the effects of BRCA1/2 variants on mRNA splicing by minigene assay. J Hum Genet 68, 65–71 (2023).
2. Zhang, R., Gao, P., Han, Y. et al. Reliable assessment of BRCA1 and BRCA2 germline variants by next-generation sequencing: a multicenter study. Breast Cancer 28, 672–683 (2021).




